The CERIT-SC Centre, an integral part of the national e-infrastructure e-INFRA CZ, has launched the second generation of AlphaFind. This fast and reliable tool for discovering similar protein structures combines massive datasets with advanced machine learning and high-performance computing power. The service is fully and freely available to the wider scientific community.
While the first generation (AlphaFindv1) enabled structure-based searching across the entire AlphaFold Protein Structure Database—accepting UniProt IDs, PDB IDs, or gene symbols—the new version builds on this concept and significantly expands it. AlphaFindv2 introduces major upgrades ranging from the search engine architecture to tools that allow researchers to work more effectively with the results.
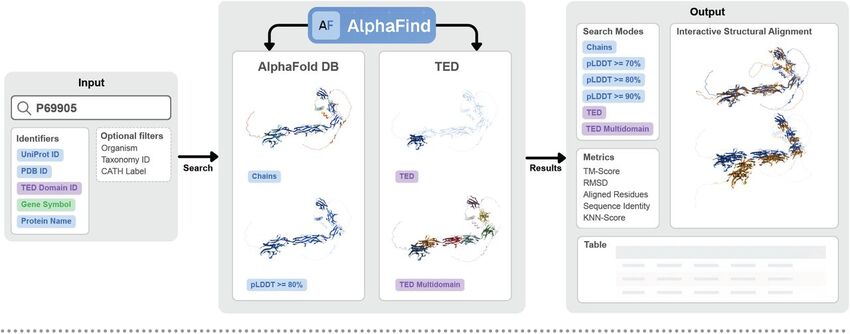
What’s new in AlphaFindv2?
- Embedding-based index. Each protein (or domain) is converted into a short numerical vector called an "embedding". By storing these vectors in an OpenSearch vector database, k-NN queries can be performed in milliseconds, even across hundreds of millions of entries.
- Asynchronous precise alignment. Following the initial high-speed search, a precise alignment calculation using US-align and TM-Score is triggered in the background. Results are updated dynamically in the table via "progressive loading," so users do not have to wait for the entire task to complete.
- Quality filtering. Before alignment, users can filter out less reliable parts of a structure based on pLDDT thresholds (≥ 70/80/90). This allows the analysis to focus exclusively on highly confident prediction regions.
- Multi-domain aggregation. For proteins with multiple domains, matches for individual domains are merged into a single "bag-of-domains" metric, which evaluates total domain coverage and quality.
- Interactive 3D visualization. Both the input and the retrieved structures are displayed in the integrated Mol* viewer, which offers smooth rotation, zoom, and the ability to adjust the weight of individual domains in the comparison.
- Export and automation. Results can be downloaded in CSV format, shared via a permanent link, or used to launch a new analysis directly from the results table. This facilitates seamless integration into subsequent research pipelines.
Who is the service for?
The AlphaFind service is available free of charge and without registration. It is designed for a wide range of users—from structural biologists and bioinformaticians to research teams involved in drug discovery.
The service is operated by CERIT-SC as part of its long-term support for scientific applications with high computing power and data capacity requirements. The method's reliability is further confirmed by its publication in the prestigious journal Nucleic Acids Research (https://doi.org/10.1093/nar/gkag372).
More information, documentation, and the interface itself are available at alphafind.ics.muni.cz, and the detailed manual can be found at alphafind.ics.muni.cz/manual.